Genomic plasticity of mollusc pathogens: causes and consequences
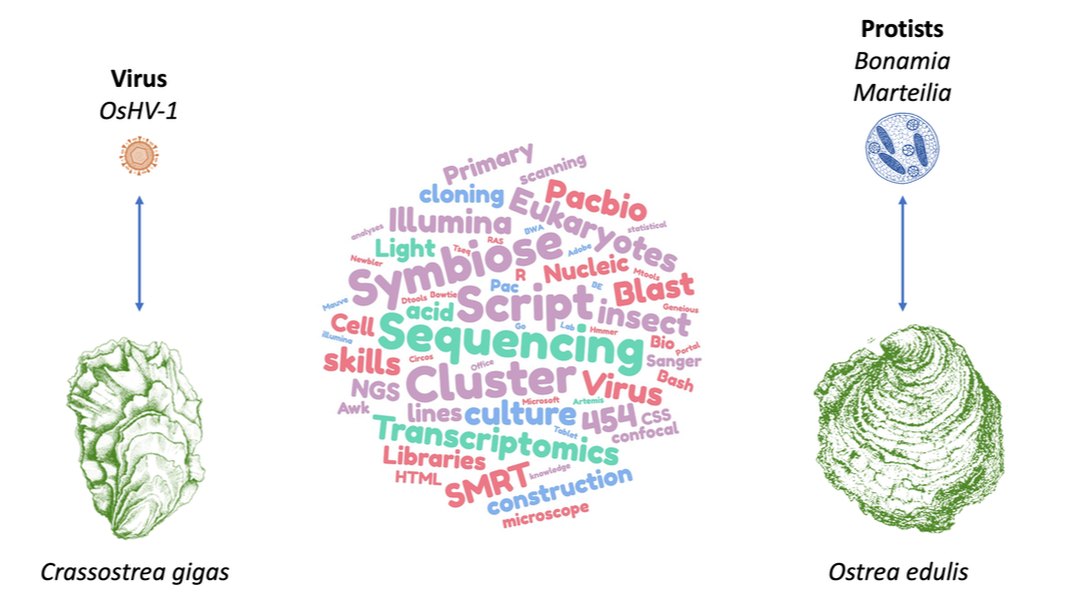
In France, oyster farming has suffered several major crises caused by pathogens. The European flat oyster Ostrea edulis practically disappeared during the 20th century during episodes of mass mortality caused by the parasites Marteilia refringens and Bonamia ostrea. In order to compensate for these losses, the Pacific oyster Crassostrea gigas was introduced in the 1970s and is now the main oyster species cultivated in France.
However, since the introduction of C. gigas, important mortality phenomena have been observed. Several factors have been identified and correlated with these mortality episodes, including: temperature, the physiological condition of the oyster, its genetic predisposition, environmental conditions and the presence of pathogens including the herpes virus OsHV-1 and opportunistic bacteria. Recently, using a holistic approach, the major steps involved in these mortality syndromes have been described. Indeed, it is the viral infection by OsHV-1 that induces a repression of the immune system of the oyster opening the way to secondary infections by opportunistic bacteria present in the microbiota of C. gigas. This syndrome is therefore well described but the evolutionary and molecular mechanisms involved are still poorly understood. The genomes of the different actors of this pathosystem are available for C. gigas, such as some variant of OsHV-1 and some of the opportunistic bacteria of the genus Vibrio. Unfortunately, the amount of genomic information currently available is insufficient to understand the genetic processes involved in such interactions and their potential evolution.
My research project proposes to use the latest advances in genome sequencing and analysis technologies to generate the data needed to understand the evolutionary processes underlying the plasticity of the genomes of infectious agents (viruses and parasites) infecting marine mollusks such as the Pacific and the flat oyster.
However, since the introduction of C. gigas, important mortality phenomena have been observed. Several factors have been identified and correlated with these mortality episodes, including: temperature, the physiological condition of the oyster, its genetic predisposition, environmental conditions and the presence of pathogens including the herpes virus OsHV-1 and opportunistic bacteria. Recently, using a holistic approach, the major steps involved in these mortality syndromes have been described. Indeed, it is the viral infection by OsHV-1 that induces a repression of the immune system of the oyster opening the way to secondary infections by opportunistic bacteria present in the microbiota of C. gigas. This syndrome is therefore well described but the evolutionary and molecular mechanisms involved are still poorly understood. The genomes of the different actors of this pathosystem are available for C. gigas, such as some variant of OsHV-1 and some of the opportunistic bacteria of the genus Vibrio. Unfortunately, the amount of genomic information currently available is insufficient to understand the genetic processes involved in such interactions and their potential evolution.
My research project proposes to use the latest advances in genome sequencing and analysis technologies to generate the data needed to understand the evolutionary processes underlying the plasticity of the genomes of infectious agents (viruses and parasites) infecting marine mollusks such as the Pacific and the flat oyster.
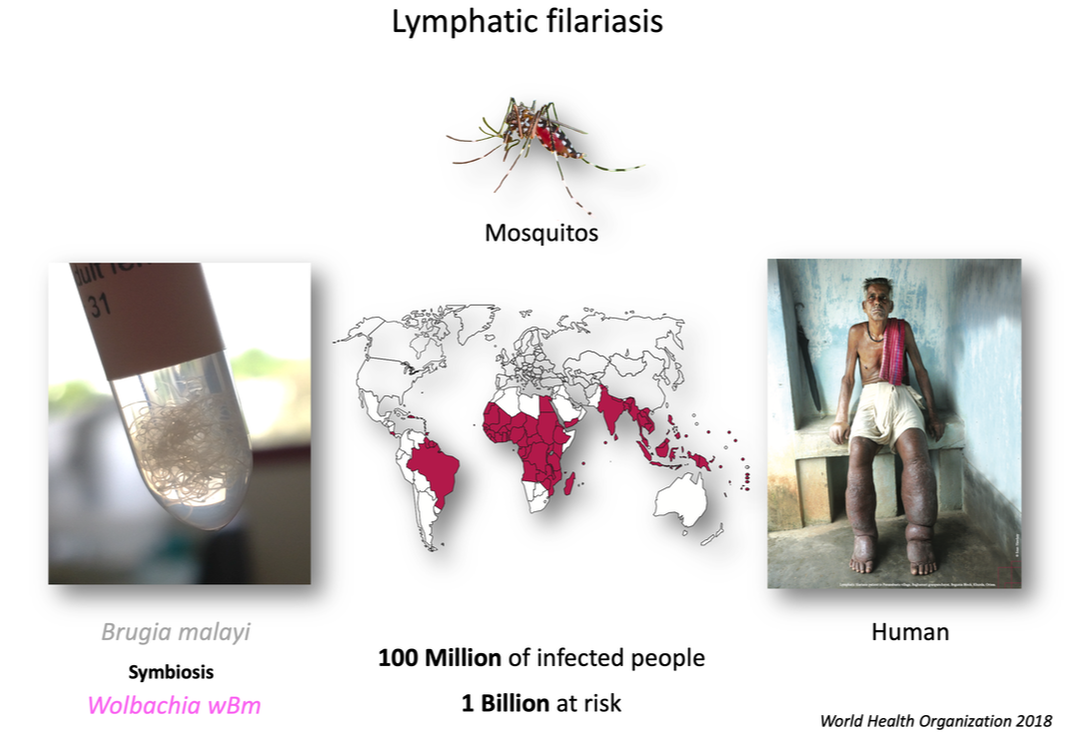
How do endosymbionts control germline stem cell maintenance? Identification and functional analyses of Wolbachia effectors in the germline of Brugia malayi, a causative agent of human filariasis
This project aimed to understand how the Wolbachia (Wb) endosymbionts control the oogenesis of parasitic filarial nematodes, responsible for debilitating human filariasis affecting 120M people.
Without Wb, filarial nematodes such as Brugia malayi become sterile, and eventually dies, making their mutualistic Wb strain (called wBm) a promising drug target to fight human and animal filariasis. The mechanisms underlying this symbiosis remain however unknown, due to two major limitations: (1) Wb are obligate intracellular bacteria, still genetically intractable, and (2) the lack of modern cell biology tools prevented so far the exploration of the parasitic nematode biology.
In this nematode sp., wBm are found in somatic hypodermal tissues in both males and females, but exclusively in the female germline. Depletion of wBm by antibiotic therapy leads to a total embryonic apoptosis. Embryos do form but fail to undergo morphogenesis. The loss of wBm impacts also the oogenesis: (i) germline proliferation is reduced, and (ii) germline stem cell (GSC) behavior and organization are greatly perturbed, suggesting a contribution from wBm to the GSC fate maintenance. In order to understand how endosymbionts control the egg production of their host and how this could affect the embryonic development, I tried to identify the germline-specific wBm effectors, and I established to what extend the host germline developmental program is dependent to the endosymbionts. To this end, I have performed a dual-RNAseq approach to study both the Wolbachia and the host nematode expression in the hypodermal and germline tissus of the female B. malayi.
Without Wb, filarial nematodes such as Brugia malayi become sterile, and eventually dies, making their mutualistic Wb strain (called wBm) a promising drug target to fight human and animal filariasis. The mechanisms underlying this symbiosis remain however unknown, due to two major limitations: (1) Wb are obligate intracellular bacteria, still genetically intractable, and (2) the lack of modern cell biology tools prevented so far the exploration of the parasitic nematode biology.
In this nematode sp., wBm are found in somatic hypodermal tissues in both males and females, but exclusively in the female germline. Depletion of wBm by antibiotic therapy leads to a total embryonic apoptosis. Embryos do form but fail to undergo morphogenesis. The loss of wBm impacts also the oogenesis: (i) germline proliferation is reduced, and (ii) germline stem cell (GSC) behavior and organization are greatly perturbed, suggesting a contribution from wBm to the GSC fate maintenance. In order to understand how endosymbionts control the egg production of their host and how this could affect the embryonic development, I tried to identify the germline-specific wBm effectors, and I established to what extend the host germline developmental program is dependent to the endosymbionts. To this end, I have performed a dual-RNAseq approach to study both the Wolbachia and the host nematode expression in the hypodermal and germline tissus of the female B. malayi.
Molecular characterization of symbiont-mediated defense of the pea aphid Acyrthosiphon pisum against the parasitoid wasp Aphidius ervi
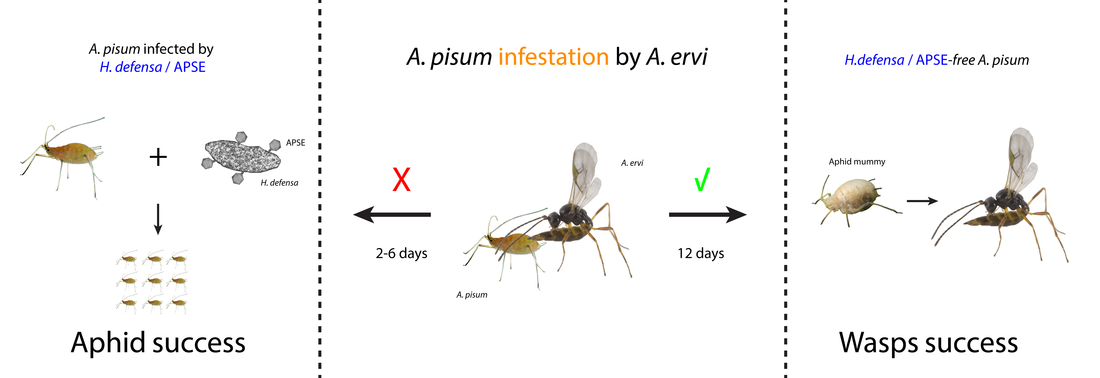
Hamiltonella defensa is a heritable secondary endosymbiont of the pea aphid, Acyrthosiphon pisum, which confers resistance to the parasitoid wasp Aphidus ervi. We know now that Hamiltonella-based protection requires bacteriophages named APSEs. Different strains of H. defensa harboring different haplotypes of APSE also give distinctive levels of protection against parasitism. In order to understand these phenotypic differences more genotypic data were needed. Because only one genome of H. defensa was available.
As postdoctoral research associate in the Strand's lab, I helped to develop an in-vitro methods for culturing H. defensa variants. Thanks to our ability to cultivate this secondary endosymbiont in-vitro, I have used the state of the art of sequencing technologies in order to perform comparative genomics and transcriptomic analyses of different strains of H. defensa. That allowed me to provide insights into the evolutionary history of H. defensa strains and shed light on H. defensa phylogeny, genome composition, and organization.
At the same time I have characterized new haplotypes of APSEs at the molecular and genomic levels.
As postdoctoral research associate in the Strand's lab, I helped to develop an in-vitro methods for culturing H. defensa variants. Thanks to our ability to cultivate this secondary endosymbiont in-vitro, I have used the state of the art of sequencing technologies in order to perform comparative genomics and transcriptomic analyses of different strains of H. defensa. That allowed me to provide insights into the evolutionary history of H. defensa strains and shed light on H. defensa phylogeny, genome composition, and organization.
At the same time I have characterized new haplotypes of APSEs at the molecular and genomic levels.
Molecular and physiological impact of the parasitoid wasp Cotesia congregata and its Polydnavirus on the insect host Manduca sexta
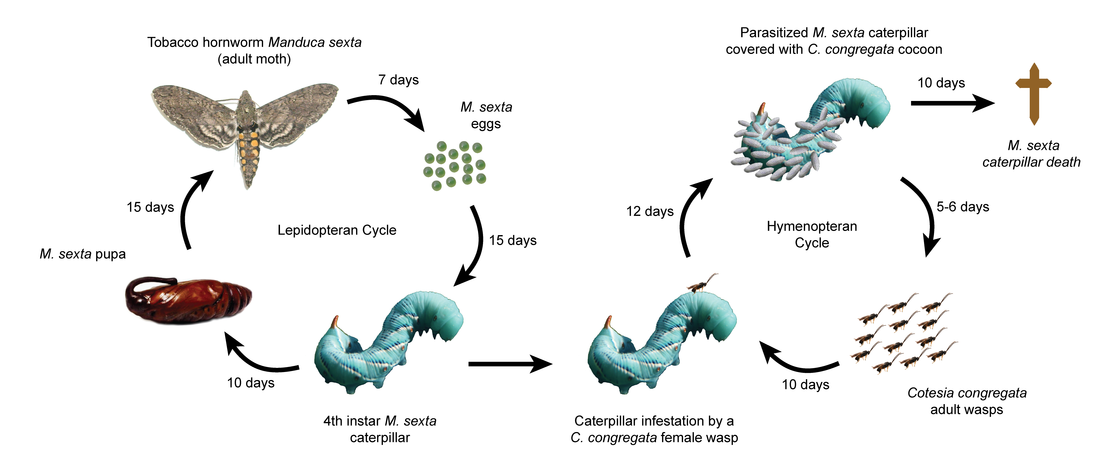
Cotesia congregata develops as a gregarious endoparasitoid inside larvae of the tobacco hornworm Manduca sexta. The parasitoid wasp has evolved virulence strategies using an obligatory viral symbiont from the Polydnavirus (PDV) family named Cotesia congregata bracovirus (CcBV). CcBV particles, containing multiple double stranded DNA circles, are produced by specialized cells of the wasp ovaries and are injected along with the eggs into the host body. They act by manipulating host immune defenses and development, thereby enabling wasps to survive in a potentially harmful environment.
The aim of my thesis was to characterize the molecular dialogue during the interaction between C. congregata and M. sexta by a high-throughput transcriptomic approach (454 sequencing technique) and by a targeted physiological approach.
Firstly, I obtained the transcriptome of two immune tissues (fat body and hemocytes) from 24 h parasitized caterpillars and via an in-depth bioinformatics analysis, I could obtain the first functional map of a PDV genome and visualize most of the M. sexta genes regulated during parasitism. The 88 expressed viral genes showed differential expression levels and I could determin that these differences in expression could be explained by the specific organization of the CcBV genome. Furthermore, fine mapping of expressed regions, confirmed experimentally that most of the viral genes contain introns, suggesting that these virulence genes have in fact a eukaryotic origin. The study of M. sexta immune gene regulation during the interaction revealed that parasitism does not affect the different pathways of the immune response in the same way. Indeed, parasitism does not prevent the activation of the antimicrobial peptide signalling pathways, suggesting that the development of the wasp eggs and larvae is occurring in an antiseptic environment. In contrast, many genes whose products are involved in (1) recognition of foreign objects, (2) humoral responses leading to phenol oxidase activation and (3) cellular immune response, are down-regulated during the interaction. This regulation is consistent with the observed inhibition of the caterpillar encapsulation defence response that involves enclosing parasitoid eggs with immune cells, followed by melanization.
Secondly, the study of the impact of parasitism on different immune cells of the host, via microscopic observations and specific markers, revealed a significant reduction in the number of adherent cells during the interaction, and the induction of a intermediate phenotype between apoptosis and pyroptosis on a category of immune cells.
My thesis work has allowed to highlight the viral genes involved in the hostparasitoid interaction and to propose a hypothesis concerning their origin. This work has also allowed to identify host targets of parasitism, notably in the pathways leading to the melanisation response and globally in cellular responses.
The aim of my thesis was to characterize the molecular dialogue during the interaction between C. congregata and M. sexta by a high-throughput transcriptomic approach (454 sequencing technique) and by a targeted physiological approach.
Firstly, I obtained the transcriptome of two immune tissues (fat body and hemocytes) from 24 h parasitized caterpillars and via an in-depth bioinformatics analysis, I could obtain the first functional map of a PDV genome and visualize most of the M. sexta genes regulated during parasitism. The 88 expressed viral genes showed differential expression levels and I could determin that these differences in expression could be explained by the specific organization of the CcBV genome. Furthermore, fine mapping of expressed regions, confirmed experimentally that most of the viral genes contain introns, suggesting that these virulence genes have in fact a eukaryotic origin. The study of M. sexta immune gene regulation during the interaction revealed that parasitism does not affect the different pathways of the immune response in the same way. Indeed, parasitism does not prevent the activation of the antimicrobial peptide signalling pathways, suggesting that the development of the wasp eggs and larvae is occurring in an antiseptic environment. In contrast, many genes whose products are involved in (1) recognition of foreign objects, (2) humoral responses leading to phenol oxidase activation and (3) cellular immune response, are down-regulated during the interaction. This regulation is consistent with the observed inhibition of the caterpillar encapsulation defence response that involves enclosing parasitoid eggs with immune cells, followed by melanization.
Secondly, the study of the impact of parasitism on different immune cells of the host, via microscopic observations and specific markers, revealed a significant reduction in the number of adherent cells during the interaction, and the induction of a intermediate phenotype between apoptosis and pyroptosis on a category of immune cells.
My thesis work has allowed to highlight the viral genes involved in the hostparasitoid interaction and to propose a hypothesis concerning their origin. This work has also allowed to identify host targets of parasitism, notably in the pathways leading to the melanisation response and globally in cellular responses.